Name:LI Jianqiang SANG Tao
Tell:
Email:lijq@wbgcas.cn sang@ibcas.ac.cn
Organization:Wuhan Botanical Garden
Researchers Reveal Population Transcriptomic Characterization of the Genetic and Expression Variation of M. lutarioriparius
2017-09-04
Miscanthus lutarioriparius is endemic to central China and found on seasonally flooded river banks, farm lands and ruderal land in cities. The species could be developed as a bioenergy resource due to its considerable biomass, presumably adapted to marginal lands, and abundant genetic variation found in natural populations. The roots of the species mainly distributed in relatively shallower soil, thus it could provide positive effects for soil and water conservation.
Supervised by Prof. LI Jianqiang & SANG Tao, YAN Juan from Wuhan Botanical Garden studied the population transcriptome of M. lutarioriparius by using 10 excellent populations with strong adaptability and high level of genetic variation.
Tests of population divergence using the transcriptome data were more sensitive using gene expression (FPKM) data. For instance, based on FPKM data, genes involved in photosynthetic processes and responses to temperature or reactive oxygen species stimuli were significantly enriched in certain populations. Additionally, nucleotide diversity was significantly negatively correlated with expression diversity within populations, while this correlation was significantly positive among populations, which suggests that genetic and expression variation play separate roles in adaptation and population persistence.
Using transcriptome data can capture faint signals of both genetic variation and expression variation, and provide a broad picture of a species’ genomic response to environmental conditions. Totally, integrating genetic and gene expression variation to study the population genetics of wild species represents a promising approach, and may uncover both adaptive and non-adaptive processes.
The work was supported by grants from the Project for Autonomous Deployment of the Wuhan Botanical Garden and the National Natural Science Foundation of China. The study entitled “Population transcriptomic characterization of the genetic and expression variation of a candidate progenitor ofMiscanthusenergy crops” was published in Molecular Ecology (DOI: 10.1111/mec.14338)
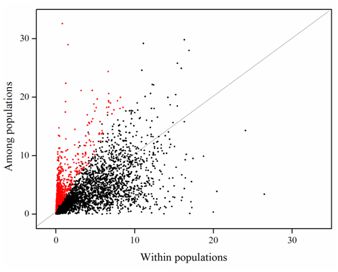
Distribution ofexpression variation within populations and among populations based on FPKM (Image by YAN Juan)