Name:Tao Shi
Tell:
Email:shitao323@wbgcas.cn
Organization:Wuhan Botanical Garden
Researchers Reveal the Integrative Expression Network of miRNA and Gene Isoforms in Lotus
2020-09-15
In plants, micro-RNAs (miRNAs) are a class of small endogenous single-stranded noncoding RNAs ranging from 18 to 24 nucleotides. They play key roles in response to plant development, abiotic and biotic stressed through regulating their target genes. RNA alternative splicing (AS) is another important post-transcriptional regulation mechanism, producing diverse transcript isoforms encoded by the same genes. The structure variation in transcript isoforms can often result in proteins with altered physical characteristics and molecular functions. In some cases, the presence or absence of the miRNA binding site in the isoform determines the possibility of its silencing by a complementary miRNA, allowing some isoforms to escape from being targeted due to lack of the miRNA binding site. The influence of miRNAs on the co-expression network of different splicing isoforms remained largely unclear.
ZHANG Yue, the doctor from CAS Key Laboratory of Aquatic Botany and Watershed Ecology, Wuhan Botanical Garden, combined the previous isoform pleomorphic result and six miRNA datasets in different developmental stages to conduct the expression network analysis of miRNA and gene isoforms in sacred lotus.
Most miRNAs had tissue-specificity in expression and a negative correlation with the expression of their targeted genes.The result of weight gene co-expression network analysis (WGCNA) revealed that preferential interactions between miRNAs and hub gene isoforms in one co-expression module was highly correlated with leaf. Intriguingly, for many genes, their corresponding isoforms were assigned to different co-expressed modules, and they exhibited that more divergence for many isoforms was escalated by both structural and expression divergence.
This study on the interactions between the miRNAs and isoforms, and functional divergence of isoforms can facilitate our current understanding of the complexity of gene regulation in plants.
This article entitled “Integrative expression network analysis of microRNA and gene isoforms in sacred lotus” is now published in BMC Genomics.
This project was supported by grants from the Strategic Priority Research Program of Chinese Academy of Sciences, the National Natural Science Foundation of China, Youth Innovation Promotion Association of Chinese Academy of Sciences, Hubei Provincial Natural Science Foundation of China, and Hubei Chenguang Talented Youth Development Foundation.
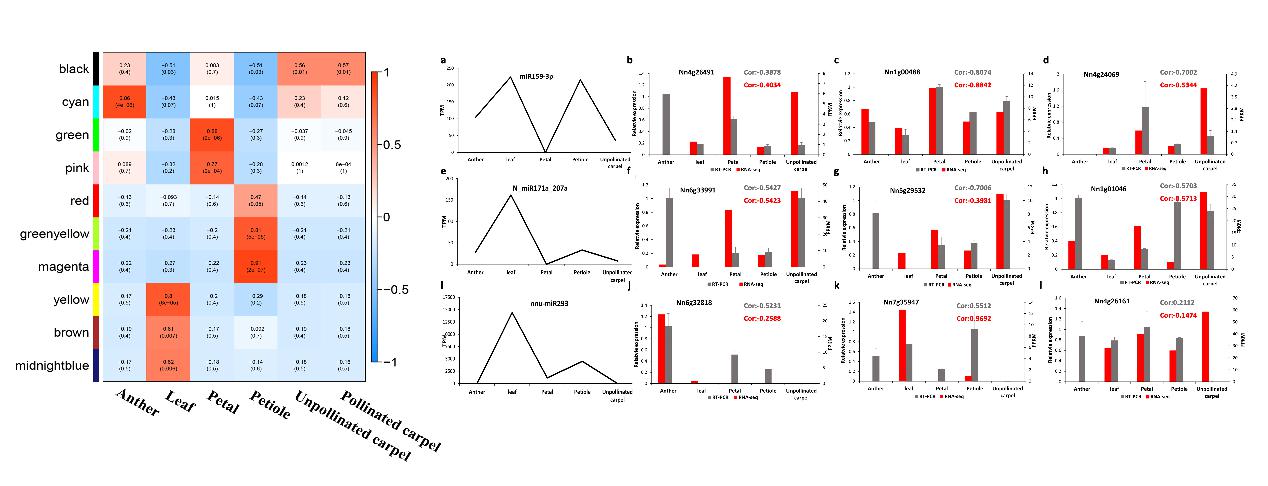
Isoform co-expression network and the expression profile of several miRNAs and targeted genes (Image by ZHANG Yue)