Name:Chen Jinming Shi Tao
Tell:
Email:jmchen@wbgcas.cn shitao323@wbgcas.cn
Organization:Wuhan Botanical Garden
Researchers Construct the Nelumbo Genome Database
2021-04-08
Sacred lotus (Nelumbo nucifera, or lotus) is an ancient aquatic dicotyledonous plant with important value in terms of understanding the origin and evolution of eudicots. It is also an important horticultural plant, and has been used as a significant ingredient for traditional medicines and food.
Researchers from Wuhan Botanical Garden and Wuhan Institute of Landscape Architecture constructed an integrative database: the Nelumbo Genome Database (NGD, http://nelumbo.biocloud.net).
The NGD deploys the latest chromosome-level genome assembly and annotations of ‘China Antique’, and it houses the sequences of 150,589 unique transcripts and 34,481 genes with complete open reading frames, as well as the sequence data of lotus gene families, transposable elements and other repeats, which providing a key foundation for deciphering the molecular genetic mechanism of sacred lotus.
Then, the NGD integrates information concerning gene expression levels and highly coexpressed genes from data from a coexpression network (WGCNA) based on 69 RNA-seq samples from 11 lotus tissues, and the cooperatively expressed gene modules, covering the developmental regulation of different organs (tissues), providing a significant public analysis platform for understanding the regulation of lotus gene expression.
Moreover, the database digitized the phenotypic traits of 88 representative cultivars, while 26,939,834 high-quality SNPs and 4,177,974 InDels from them are deployed in the NGD, which enriches the genetic variation information of lotus.
Furthermore, the NGD database deploys multiple open source and free bioinformatics tools including Blast, Gbrowse, Primer Design, and Expression Visualization, and these accessible applications provide users with convenience in sequence analysis, genomic mutation search and visualization, highly co-expressed gene network construction and visualization, variety trait query, etc.
In conclusion, the lotus public data platform offers valuable reference information for investigation and experimental design in the fields of lotus genome evolution, molecular genetics, and molecular breeding.
This study was financially supported by grants from the CAS Strategic Priority Research Program, the National Natural Science Foundation of China, the Youth Innovation Promotion Association of the CAS, the Bureau of Landscaping and Forestry of Wuhan Municipality, the Hubei Provincial Natural Science Foundation of China, and the Hubei Chenguang Talented Youth Development Foundation. Thanks for Razgar Seyed Rahmani from Ghent University for the discussion.
These findings were published online in Scientific Data entitled "Nelumbo genome database, an integrative resource for gene expression and variants ofNelumbo nucifera".
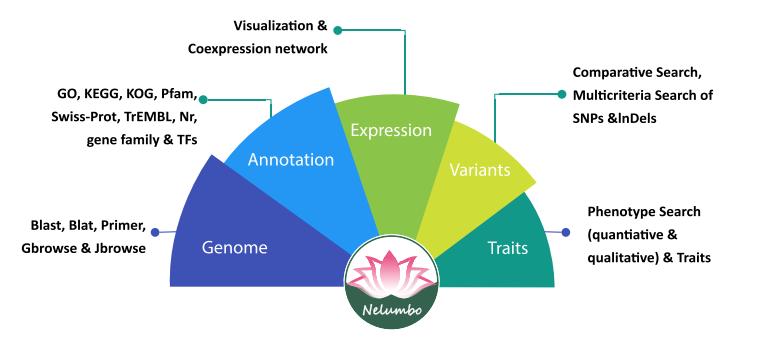
A schematic of the data collection and utilities for the Nelumbo Genome Database (Image by LI Hui)